News Spotlight
Tuesday, October 22, 2024
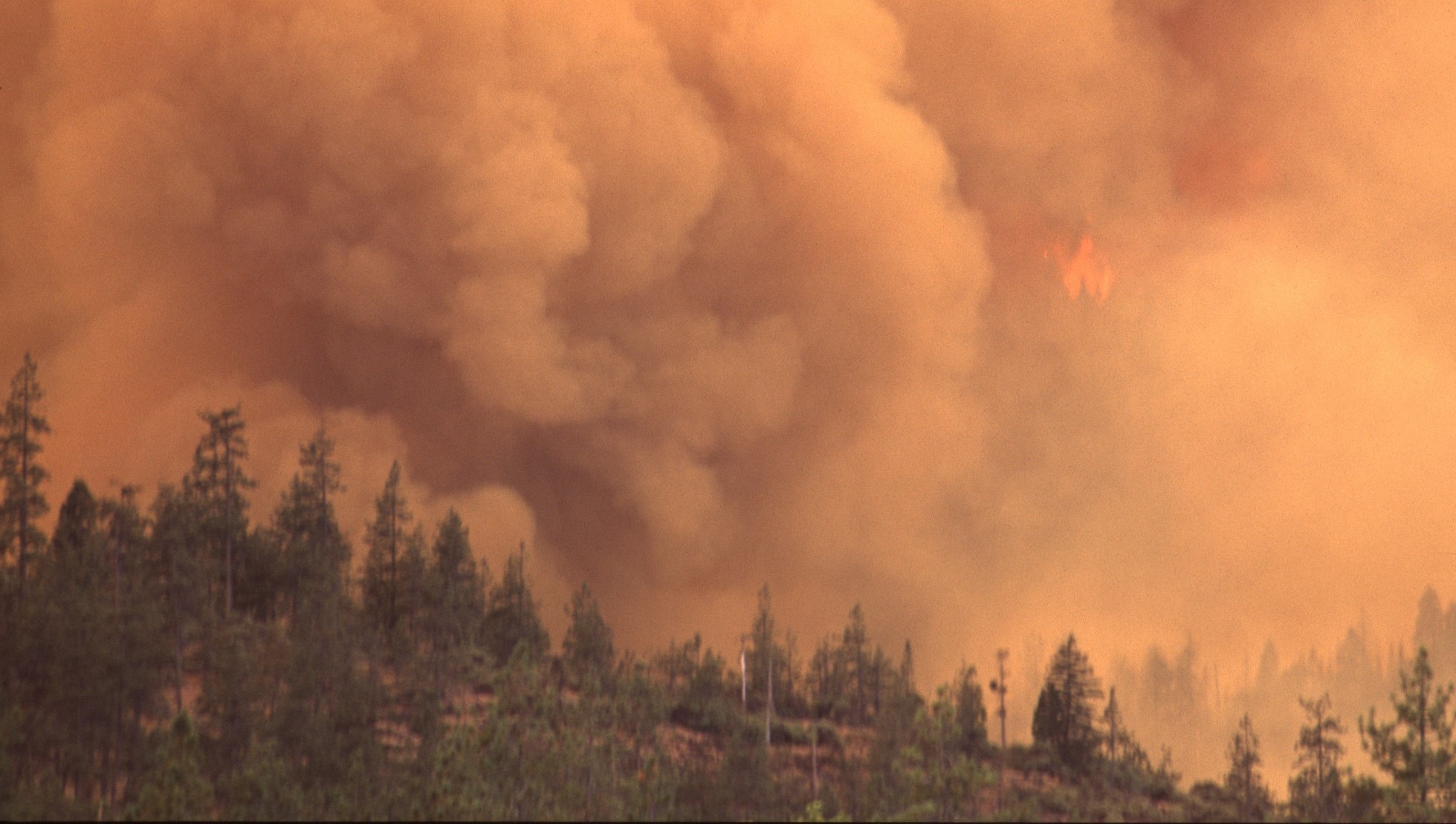
Wildfires Alter Air Pollution Patterns Across North America
Researchers from the National Center for Atmospheric Research use Supercomputer to study impact of wildfires on air pollution in North America
See Past Spotlights >
©1994-2024
|
Shodor
|
Privacy Policy
|
NSDL
|
XSEDE
|
Blue Waters
|
ACM SIGHPC
|
 |
|
 |
|
 |
|
 |
|
 |
XSEDE Code of Conduct
|
Not Logged In. Login
|
XSEDE Code of Conduct
|
Not Logged In. Login
 |
|
 |
|
 |
|
 |
|
 |
XSEDE Code of Conduct
|
Not Logged In. Login
|
XSEDE Code of Conduct
|
Not Logged In. Login